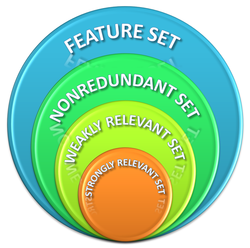
Feature (Attribute) Selection
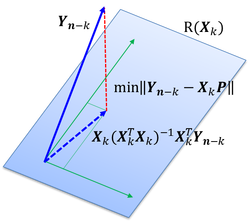

Unsupervised Feature Selection Using Incremental Least Squares: The Least Square Error (LSE) of approximating the complete data set via a reduced feature subset was proposed as the quality measure for feature selection. Guided by minimization of the LSE, a kernel least squares forward selection algorithm (KLS-FS) was developed that is capable of both linear and non-linear feature selection. An incremental LSE computation was designed to accelerate the selection process and, therefore, enhances the scalability of KLS-FS to high-dimensional data. The proposed method was implemented and included in the RapidMiner extension (plug-in) available on the software page.

Spatial Distance Join based Feature Selection: A Spatial Distance Join (SDJ) based feature selection method (SDJ-FS) was developed, which extended the concept of Correlation Fractal Dimension (CFD) to handle both feature relevance and redundancy jointly for supervised feature selection problems. The Pair-count Exponents (PCEs) for the SDJ between different classes and that of the entire dataset (i.e., the CFD of the dataset) are proposed respectively as feature relevance and redundancy measures. For the SDJ-FS method, an efficient divide-count approach of backward elimination property is designed for the calculation of the SDJ based feature quality (relevance and redundancy) measures. The proposed method was implemented and included in the RapidMiner extension (plug-in) available on the software page.
Development of (Q)SARs for Engineered Nanomaterials
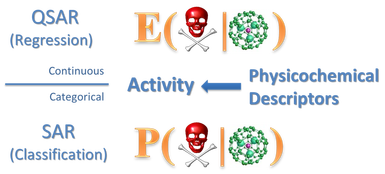
(Q)SAR can be considered as an application of data mining to the fields of chemical, material and biological sciences, and engineering. It is essentially a classification/regression model that is develeped to predict (bio-)activity of chemicals/nanomaterials based on their physicochemical (structural) properties. The assumption for (Q)SARs is that chemicals of similar physicochemical properties are likely to have similar (bio-)activity.
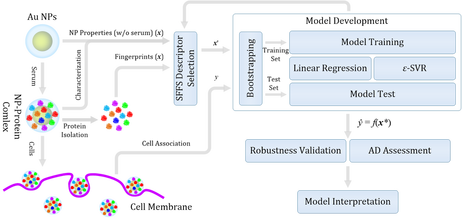
QSARs for Cellular Association and Uptake of Nanoparticles: Both linear and non-linear support vector regression (ε-SVR) models were used along with sequential forward floating descriptor (feature) selection for the QSAR development. For the ε-SVR model, the regularization factor (C), tube size (ε), and kernel width (γ) were determined according to a recommended practical model parameter selection approach. Two sets of the QSARs were developed using the above approach. One was for predicting the cellular association of Au nanoparticles (NPs) using their corona proteins and physicochemical properties while the other one was for cellular uptake of NPs of the same iron oxide core but with different surface-modifying organic molecules.
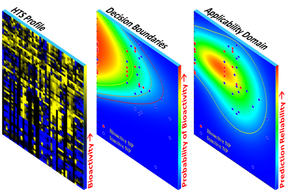
Classification (Q)SAR for Bioactivity of Iron Oxide Core NPs: A classification (Q)SAR was developed using a set of bioactivity profiles for four cell types and four different assays. The developed (Q)SAR was as Naive Bayesian model with spin-lattice relaxivity and zeta potential selected as its descriptors. The establishment of a quantitative (Q)SAR applicability domain was demonstrated, making use of a probability density. In addition, decision boundaries were determined for the above (Q)SAR for different acceptance levels of false negative to false positive predictions.
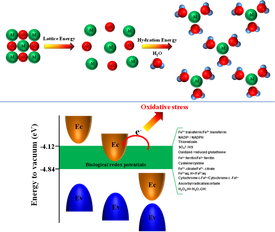
Classification (Q)SARs for Metal Oxide NP Toxicity: A range of models (e.g., Naive Bayesian classifier, linear regression, linear discriminate analysis, logistic regression, quadratic logistic regression, and SVM) were evaluated, based on an initial pool of thirty NP descriptors for Metal Oxide NPs. The conduction band energy and ionic index (often correlated with the hydration enthalpy) were identified as suitable NP descriptors that are consistent with suggested toxicity mechanisms for metal oxide NPs and metal ions. The best performing (Q)SAR was a SVM model with the above two descriptors arrived at a high balanced classification accuracy.
Nanoinformatics
Nanoinformatics that extracts, validates, stores, shares, analyzes, models, and applies information pertaining to nanotoxicity data is necessary in decision making for ENM design, optimize, manufacturing & processing, exposure control, as well as Environmental, Health, and Safety (EHS).
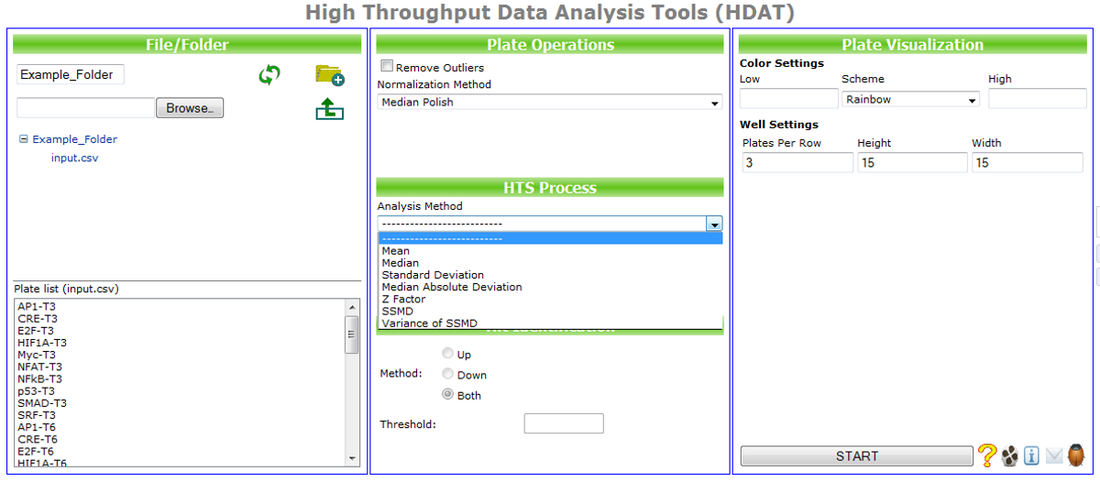
As part of the effort for build nanoinformatics infrastructure, we developed an integrated set of High Throughput Data Analysis Tools (HDAT) and made it available through UC CEIN's nanoinformatics website for public access. HDAT is an online HTS data analysis tool that provides: 1) a set of functions for HTS plate data pre-processing and analysis tools for Outlier Removal, Data Normalization, Data Summarization, Hit Identification, Data Visualization (Self-Organizing Map and Heatmap), and Clustering Analysis, 2) a standard and reference platform for analyses and comparisons across different HTS experiments, and 3) a standardized HDAT plate data format with flexible configuration for different matrixed data.
As part of the effort for build nanoinformatics infrastructure, we developed an integrated set of High Throughput Data Analysis Tools (HDAT) and made it available through UC CEIN's nanoinformatics website for public access. HDAT is an online HTS data analysis tool that provides: 1) a set of functions for HTS plate data pre-processing and analysis tools for Outlier Removal, Data Normalization, Data Summarization, Hit Identification, Data Visualization (Self-Organizing Map and Heatmap), and Clustering Analysis, 2) a standard and reference platform for analyses and comparisons across different HTS experiments, and 3) a standardized HDAT plate data format with flexible configuration for different matrixed data.
|
In one of my recent studies, an unsupervised feature selection method was developed for analysis of data sets of high dimensionality. The Least Square Error (LSE) of approximating the complete data set via a reduced feature subset was proposed as the quality measure for feature selection. Guided by minimization of the LSE, a kernel least squares forward selection algorithm (KLS-FS) was developed that is capable of both linear and non-linear feature selection. An incremental LSE computation was designed to accelerate the selection process and, therefore, enhances the scalability of KLS-FS to high-dimensional datasets. The proposed method was implemented as a rapidminer 5 extension (plug-in), which is available for downloading under the software page of this site.
|